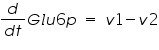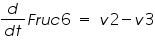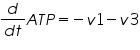| . | . | |||
| . | Import and simulate models from different databases |
. | ||
| . |
Objective:
Introduction:
Systems biology is concerned with modeling of biological systems at a systems level. Consider the interactions of all the components of a system rather than its isolated properties. Biological systems such as cells, regulatory gene networks and protein interaction complexes cannot be understood from individual components like proteins, genes and mRNA. It must be understood through the networks involving all components at the same time. It is mainly focusing on the stochastic modeling and computer simulation techniques for analyzing the systems.
Biomodel is not just a repository it is a true database of computational models of biological systems that have been described in the peer-reviewed scientific publications. This repository (database) allows one to store, search, and retrieve the mathematical models. These models are curated with the external resources like publications, ontologies, databases of compounds and pathways. The models and their related information is stored in the form of MySQL tables. This allows one to search not only the particular models based on their internal components but also based on the extensive additional annotation.
JWS Online is a tool for simulation of kinetic models from a database. Hence JWS online is a repository for curated biochemical pathways and simulation tool for these models in CellDesigner. It is easy to download the xml format from the database directly. By clicking the ModelDatabase tab to access the models, or the other tabs for more information. User can easily download and view the model from database.
Biomodel repository (database) allows one to store, search, and retrieve the mathematical models .By typing the model name in the query box given in Bomodel.net database. it can redirect to the page where we can download the model in required format. Home page of the database is shown below. Biomodel database relies on two internal databases like Biomodels and web-auth.
Figure 1 : Home page of Biomodels.net database.
JWS Online is a tool for simulation of kinetic models from a curated database. By clicking the Model Database tab to access the models, or the other tabs for more information. In the list of models, there is a run button for each model. Clicking on this it will redirects to the model selection to the server, which generates and returns a java applet. The window is divided into three frames. In the left frame, the user can modify the parameters. The top right frame shows a representation for the reactions in the model. The right side of this frame contains three tabs, allowing a time course simulation, steady state calculations. To run the simulation, the user can choose the start and end times.
Figue 2: JWS online databse home page
What is a model:
A model is a representation of any object or a system .It is important to highlight that a model is not the real object, but only a human construct to help us to understand real object systems. In general all models have an input, a processor, and an output of expected results. In mathematical modelling, we translate those values into the language of mathematics.
What is a mathematical model :
We want to know how to make or generate or model the mathematical representation .Once the structure of a model has been determined, the mathematical equations must be chosen to describe the system. It is worth to choose these formulation carefully because they may have unexpected effects on the behaviour of the model. The modeling part mainly depends on the formulation of the model,analyse the model, interpret it and test the model.
Process of mathematical modelling :
Figure 3: Process of Mathematical modeling
Steps involved in Mathematical modelling :
Mathematical modeling and simulation :
The metabolic network consists of reactions where one type of molecule is transformed to another type. A simple mathematical equation for a metabolic network is as follows.
Figure 5: Simple network
The ODE system for this reaction system is given by:
[v = k * conc.of the reactants]
Where: v= Rate of reaction; k= Rate constant. Calculation:
NOTE: The rate of change of Glucose-6-phosphate is the rate of formation of Glucose-6-phosphate (v1) and the rate of degradation of Glucose-6-phosphate (-v2).
Here : -v represents degradation.
Rate of change of Glucose-6-phosphate = rate of formation of Glucose-6-phosphate (v1)- rate of degradation of Glucose-6-phosphate (-v2,-v3)
CellDesigner :
Systems biology is deals with mainly modeling of biological systems at system level. The main purpose of CellDesigner is to draw networks using the symbolic notation, this system is proposed by Kitano. It is easy to retrieve the model from the databases.
Features of cell designer :
It is easy to make a model or to built a network based on mathematical equations.
Repository (database) allows one to store, search, and retrieve the mathematical models. These models are curated with the external resources like publications, ontologies, databases of compounds and pathways.
Different format is needed for computing a model to the point where it can be simulated and analyzed. Where the Systems Biology Markup Language (SBML) comes in this analization.
Select a particular model and run the simulation by changing the parameters. The results can be viewed graphically with the time on the X-axis and species (or substances) on the Y-axis. It implies that, what will be expression of particular substance at a particular time, based on an equation.
Cite this Simulator: |
..... | ||
| ..... | ..... | |||
 |
Copyright @ 2024 Under the NME ICT initiative of MHRD |
 |